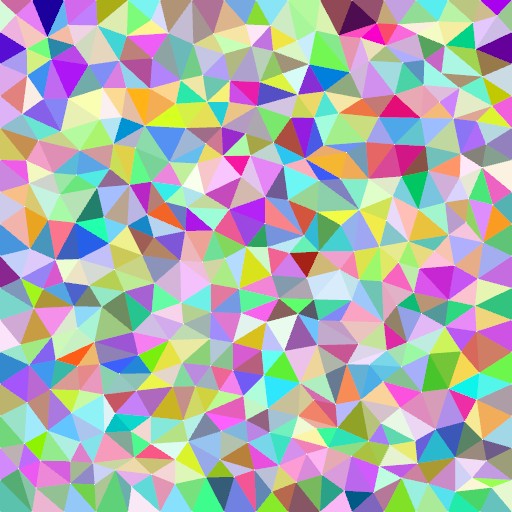


Some computer graphics applications require random, yet somehow regular meshes – examples include material simulation or surface reconstruction.
Poisson disc sampling is excellent for creating a vertex field with tight upper and lower limits on the distance between neighboring vertices (a.k.a. 2D blue noise). Such a sampling also guarantees that, on average, every vertex has six neighbors.
But how to connect these vertices? Requirements normally include:
islands
This is usually solved with Delauney triangulation. In general, 2D Delauney triangulation has a runtime of
If you use a grid to accelerate the generation of the 2D Poisson distribution, you can recycle this grid to speed up the triangulation. It won’t be perfect, but the algorithm is cache-friendly and extremely fast. It runs in
It runs a scan line through the grid and adds triangles to the graph as it encounters new vertices. Both the total number of triangles and the maximal number of vertices in the scan line are limited by properties of the distribution, so no dynamic memory allocation needs to take place in the inner loop.

X points right and Y points up. Triangles are defined in clockwise order.
The Poisson distribution must be created using a 2D grid as explained in Fast Poisson Disk Sampling in Arbitrary Dimensions. You want to do this anyway because it generates points in
The distance between neighbors must be in
Unlike Delauney triangulation, this algorithm will not always generate optimal triangulation. In my tests, 2 % of triangle edges turned out longer than

Don’t use this algorithm if this is a problem for you!
There should be samples at the outer bounds of the field or the edges will turn out horribly deformed. Note that this same limitation applies to constrained Delauney. The images in this article place samples in regular intervals at the outer edges.
There must be at least two lines and two columns in the grid.
Start at
Note that the total number of triangles in the graph is limited to six times the number of samples. Reserve this, too, and there will be no memory allocation in the inner loop.
Iterate all grid lines towards positive Y. Iterate each grid line towards positive X. If you encounter a vertex, find the vertex in the wavefront just below and left to it by iterating the wavefront until
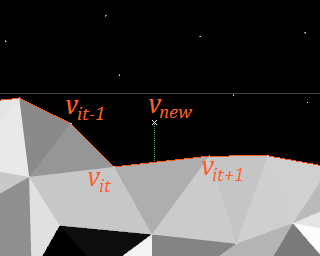
Now either add a new vertex to the wavefront or replace an existing one. In doing so, it will create either one or two triangles.
In the best case,
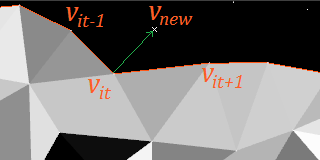
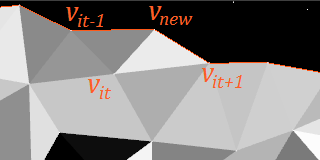
You should, in particular, check both distances
Note that the wavefront must satisfy several requirements for this to be legal:
If the above is not possible, try extending the wavefront by a new vertex:
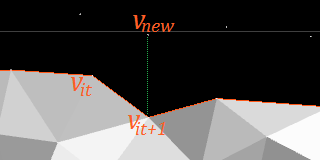
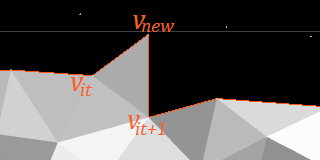
After traversing a grid line, the wavefront may have more vertices than before. This is a dangerous trend that, if continued, would totally fragement the wavefront.

If there is a cavity – a vertex that is farther than
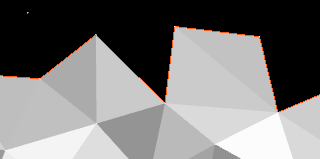
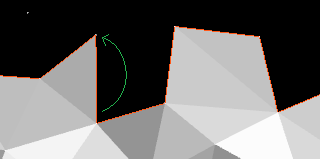

I found that triangulation quality improves if you also remove any concave vertices, given the lengths of the new triangle edges are all below
The factor of
Wavefront cleanup could be combined with wavefront operations above (specifically, cleanup could run while the wavefront is traversed to find the segment just below a new sample), but I didn’t bother doing it yet.
The algorithm above cannot properly handle the edges because the leftmost vertex of the wavefront is never replaced. If you need the left and right edges properly triangulated: Check if the first and last grid cells contain vertices. If so, replace the first and last vertex of the wavefront, creating one new triangle, respectively
// Initialization create_poisson_2d(points, w, h, grid, wgrid, hgrid) // Initialize the wavefront with the first grid line: list_of_indices wavefront; for vertex_index in grid.lines[0] wavefront.add(vertex_index) for line in grid.lines(1...hgrid) // If this line starts with a new point, move the wavefront there. This saves us from handling special cases later on. if is_set(line[0]) add_triangle(wavefront[0], line[0], wavefront[1]) wavefront[0] = line[0] // Process all vertices in this line: wavefront_it = &wavefront[0] for x in 1...wgrid if not is_set line[x] continue; // Found a vertex. Find the beginning of the corresponding line segment in the wavefront. new_vertex_index = line[x] while points[wavefront_it[1].x < points[new_vertex_index].x ++wavefront_it; // Decide whether to move an existing vertex or to create a new one. if wavefront_it >= &wavefront[1] && points[wavefront_it[-1]].y > points[wavefront_it[0]].y // left vertex is lower than its left neighbor && dist(points[wavefront_it[-1]], newVertex) < 2 * r && dist(points[wavefront_it[1]], newVertex) < 2 * r if dist(points[wavefront_it[0]], newVertex) < dist(points[wavefront_it[-1]], points[wavefront_it[1]]) add_triangle(wavefront_it[-1], new_vertex_index, wavefront_it[0]) add_triangle(wavefront_it[0], new_vertex_index, wavefront_it[1]) else add_triangle(wavefront_it[-1], new_vertex_index, wavefront_it[1]) add_triangle(wavefront_it[-1], wavefront_it[1], wavefront_it[0]) wavefront_it[0] = new_vertex_index; else // create new vertex add_triangle(wavefront_it[0], new_vertex_index, wavefront_it[1]) wavefront.remove(wavefront_it + 1) wavefront_it[1] = new_vertex_index // If this line ends with a new point, move the wavefront there. This saves us from handling special cases above. if is_set line[wgrid - 1] add_triangle(line[wgrid - 1], wavefront[wavefront.length - 1], wavefront[wavefront.length - 2]) wavefront[wavefront.length - 1] = line[wgrid - 1] // Fill any cavities in the wavefront. wavefront_it = &wavefront[0] final_y = y / hgrid * h - 1.28f * r // any point below this coordinate was not triangulated correctly for wavefront_it in &wavefront[0]...&wavefront[wavefront.length - 2] is_stretched = dist(points[wavefront_it[0]], points[wavefront_it[2]]) > 2 * r is_concave = signedArea(points[wavefront_it[0]], points[wavefront_it[1]], points[wavefront_it[2]]) < 0 if points[wavefront_it[1]].y < finalY add_triangle(points[wavefront_it[0]], points[wavefront_it[2]], points[wavefront_it[1]]) wavefront.remove(wavefront_it + 1)